Genetics
Abstract
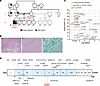
Germline BRCA1/2 pathogenic variant (PV) carriers have elevated young-onset breast cancer risk. To define the pretreatment genomic landscapes of young-onset gBRCA-associated breast cancer, we evaluated 136 treatment-naïve tumors diagnosed before age 50 (92.6% ≤40): gBRCA1 86(63.2%); gBRCA2 50(36.8%) in the prospective POSH study, and 66 noncarriers from The Cancer Genome Atlas. Using whole exome sequencing, we analyzed somatic variation, allele-specific loss of heterozygosity (asLOH), homologous recombination deficiency (HRD), and single-base substitution signatures (SBS). gBRCA1(93%) and gBRCA2(96%) breast cancers had high rates of asLOH, but differed significantly in average HRD scores (57.4 ± 1.3 vs 43.7 ± 1.5, P < 0.0001) and median SBS composition (%): SBS1 (aging-associated) 12.9 vs 7.3, P = 0.013; SBS18 (reactive oxygen species [ROS]-associated) 1.4 vs 0, P = 0.007; and SBS3 (HRD-associated) 27.3 vs 42.6, P = 0.002. Compared to gBRCA2 tumors, gBRCA1 tumors with asLOH were significantly enriched for alterations in Hallmark ROS, DNA repair, and epithelial-mesenchymal transition pathways. In ER-positive, HER2-negative tumors from gBRCA1/2 carriers compared to noncarriers, we found significant enrichment of RB1 (OR:6.3;95%CI:2.8–15.4;padj = 0.001), TP53 (OR:4.6;95%CI:1.9–12.1;padj = 0.017), FAT1 (OR:3.9;95%CI:1.84–8.7;padj = 0.013), and MYC (OR:4.0;95%CI:1.8–9.1;padj = 0.017) SNV/indels/CNVs, associated with CDK4/6i resistance. Together, these findings demonstrate significant differences between gBRCA1 and gBRCA2-associated breast cancers, and preexisting CDK4/6i resistance mechanisms supporting prospective trials with individualized therapy for gBRCA1 vs gBRCA2 carriers, and comparing PARPi to CDK4/6i for ER-positive gBRCA1/2-associated breast cancer.
Authors
Mwangala P. Akamandisa, Mingyi Xia, Wilson Cheah, Bradley Wubbenhorst, Kurt P. D'Andrea, Mengyao Fan, Jake S. Shilan, Dana Pueschl, Anupma Nayak, Hayley McKenzie, William Tapper, Ellen R. Copson, Ramsey I. Cutress, Susan M. Domchek, Diana M. Eccles, Katherine L. Nathanson
Abstract
Survival after lung transplantation is limited by chronic, progressive graft failure, termed chronic lung allograft dysfunction (CLAD). Graft-resident mesenchymal cells (MCs) drive CLAD pathogenesis and exhibit stable dysregulated signaling, yet the transcriptomic and epigenomic drivers underlying this fibrogenic transformation remain elusive. We used single-cell multi-omic profiling to characterize gene expression and chromatin accessibility in MCs isolated from lavage fluid of lung transplant recipients with and without CLAD, collected early post-transplantation or after disease onset. MCs obtained after CLAD onset demonstrated a distinct transcriptomic signature compared with non-CLAD controls, enabling classification of disease status at the single-cell level with > 98% accuracy using signature genes. Chromatin accessibility analyses identified enrichment of CCAAT-enhancer-binding protein family transcription factors, specifically CEBPD, in CLAD MCs. Early post-transplant MCs showed minimal accessibility differences, suggesting that CEBPD-associated regulatory changes emerge over time. Integration analyses identified eight MC states and a CLAD-specific shift towards a fibrotic state. CEBPD, SOX4, and FOXP2 were identified as putative regulators of this state with substantial overlap in predicted targets. Targeting CEBPD reversed fibrotic phenotypes of CLAD MCs (decreased ECM expression, contractility, proliferation, and migration). Together, these data provide insights into transcriptomic and epigenomic changes in post-transplant MCs, nominating biomarkers and therapeutic targets.
Authors
Lu Lu, A. Patrick McLinden, Natalie M. Walker, Ragini Vittal, Yichen Wang, Fatemeh Fattahi, Stephen T. Russell, Michael P. Combs, Joshua D. Welch, Vibha N. Lama
Abstract
Nearly 100 individuals have been identified who carry deleterious biallelic germline variants in CARD9 and experience life-threatening, invasive fungal infections caused by Ascomycetes but are otherwise resistant to other infectious agents. CARD9 is an adaptor protein expressed predominantly in myeloid cells, which functions downstream of dectin receptors, pattern recognition receptors for fungal antigens, to activate innate immune responses. The impact of CARD9 deficiency on lymphocytes, however, is less clear. We deciphered the functional consequences and delineated mechanisms of disease in a patient (P1) with a nonsense germline homozygous CARD9 variant (c.673A>T/p.K225*) and invasive Candida disease. P1’s PBMCs expressed truncated CARD9 and showed significantly reduced cytokine production in response to fungal ligands. P1 had reduced frequencies of circulating memory CD4+ TH17-like (CCR6+CXCR3–) cells. In addition, in vitro differentiation of P1’s naive CD4+ T cells into IL-17A/IL-17F–secreting cells was greatly impaired. Consistent with impaired responses of innate and adaptive immune cells from P1 in vitro, proportions of Candida-specific CD4+ T cells were strongly and selectively diminished. Our findings suggest that the CARD9 variant identified in P1 is pathogenic, affecting not only CARD9-induced immunity mediated by myeloid cells but also CD4+ T cell–intrinsic IL-17–dependent immunity and Candida-specific T cell responses.
Authors
Erika Della Mina, Carlos G. El-Haddad, Timothy A. West, Clara W.T. Chung, Jing Jing Li, Vivienne Lea, Elissa K. Deenick, Filomeen Haerynck, Jean-Laurent Casanova, Anne Puel, Cindy S. Ma, Stuart G. Tangye, Alisa Kane
Abstract
Prurigo nodularis (PN) is a chronic inflammatory skin disease characterized by pruritic skin nodules of unknown etiology. Little is known about genetic changes in PN pathogenesis, particularly somatic events, which are often implicated in inflammatory conditions. We thus performed whole-exome sequencing on 54 lesional and nonlesional skin biopsies from 17 patients with PN and 10 patients with atopic dermatitis (AD) for comparison. Somatic mutational analysis revealed that PN lesional skin harbors recurrent somatic mutations in fibrotic, neurotropic, and cancer-associated genes that are absent in adjacent PN nonlesional skin. Nonsynonymous mutations were most frequently present in NOTCH1 and the Notch signaling pathway, a key regulator of cellular proliferation and tissue fibrosis. In contrast, NOTCH1 mutations were absent in AD. Somatic copy-number analysis, combined with expression data, identified recurrently deleted and downregulated genes in PN lesional skin, which are associated with axonal guidance and extension. Follow-up immunofluorescence validation demonstrated increased NOTCH1 expression in PN lesional skin fibroblasts and increased Notch signaling in PN lesional dermis. Finally, a multicenter analysis revealed increased risk of NOTCH1-associated diseases in patients with PN. In characterizing the somatic landscape of PN, this study highlights the potential role of Notch pathway dysregulation in PN pathogenesis and fibrosis.
Authors
Ahmad Rajeh, Shahin Shahsavari, Hannah Cornman, Alexander Kollhoff, Anuj Gupta, Mindy D. Szeto, Anusha Kambala, Olusola O. Oladipo, Varsha Parthasarathy, Junwen Deng, Melika Marani, Shirin Shahsavari, Selina M. Yossef, Vedha Vaddaraju, Waleed Adawi, Yagiz M. Akiska, Davies M. Gage, Sarah Wheelan, Thomas Pritchard, Madan M. Kwatra, Yevgeniy R. Semenov, Alexander Gusev, Won Jin Ho, Srinivasan Yegnasubramanian, Shawn G. Kwatra
Loss of Tumor-Infiltrating Lymphocytes and Poor Response to Immunotherapy in IDH GOF Mutant Melanoma
Abstract
Recent innovations in melanoma treatment with immune checkpoint blockade (ICB) have improved overall outcomes for patients, however over 50% of patients still develop resistance to treatment. These patients either have intrinsic resistance, and never respond to therapy, or develop acquired resistance months or years into treatment. The mechanisms underlying ICB resistance remain poorly understood. Our data shows that isocitrate dehydrogenase gain of function (IDH GOF) mutant melanoma patients have a worse response to anti-PD1 immunotherapy. IDH mutations have been found to be oncogenic and associated with differential methylation in multiple cancers but are not yet characterized in human melanoma. Here, we investigate the clinical, immune, and transcriptional phenotypes of IDH GOF melanomas through analyses of clinical response, single-cell RNA sequencing, bulk RNA sequencing, and DNA methylation data. Single-cell data analysis shows decreased immune infiltrate and activity in the IDH GOF tumors. Bulk sequencing data demonstrates the association between IDH mutation, immune exclusion, and disruptions in global DNA methylation. The melanoma-derived genomic data presented supports previously described resistance mechanisms of IDH mutation in other cancer types and is the first demonstration of the role of IDH GOF in the human melanoma tumor microenvironment.
Authors
Emma Specht, Lakshmi Pakanati, Meng-Ju Wu, Russell W. Jenkins, Derek N. Effiom, Nabeel Bardeesy, Bradley E. Bernstein, Moshe Sade-Feldman, Christine G. Lian, Genevieve M. Boland, Elena Torlai Triglia, Sonia Cohen
Abstract
X-linked myotubular myopathy (XLMTM) is a rare genetic disorder that typically presents at birth with progressive muscle weakness and respiratory difficulties and is caused by myotubularin-1 (MTM1) gene mutations. Here we examine the role of phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 beta (PIK3C2B), a lipid kinase that interacts with MTM1, in XLMTM in various models. We examined the effect of BLU3797, a novel, highly potent, selective, orally bioavailable PIK3C2B inhibitor, on survival, muscle development, myofiber phenotypes, and gene expression in MTM1-/y mice. PIK3C2B-deficient XLMTM animals demonstrated increased survival, restored muscle function, fewer myofibers with centralized nuclei, and normalization of disease-associated molecular markers. BLU3797 alleviated the XLMTM phenotype in a dose-dependent and reversible manner. Loss of functional PIK3C2B in XLMTM mice promoted a more differentiated, adult-like myofiber profile, which was strongly associated with normalization of disease surrogates and a reduction in markers of early muscle development and regeneration. BLU3797 treatment appears to modulate the expression of microRNAs associated with satellite cell activation and myofiber fusion. These findings indicate that PIK3C2B inhibition with BLU3797 effectively reverses the XLMTM disease phenotype by enhancing muscle function and promoting development toward a more mature state.
Authors
Andrew Shearer, Melissa L. Brooks, Maxine M. Chen, Thiwanka Samarakoon, John Hsieh, Gramoz Kondakci, Emanuele Perola, Jason Brubaker, Kristina Fetalvero, Stefanie Schalm, Joana Caetano-Lopes
Abstract
VIC-1911 (formerly TAS-119) is a next-generation, ATP-competitive Aurora kinase A (AURKA) inhibitor with a favorable biosafety profile. However, it has not been evaluated in prostate cancer (PC), wherein AURKA is highly expressed in advanced stages and represents a critical therapeutic target. Here, we demonstrate that VIC-1911 potently inhibits AURKA activity with high selectivity over AURKB/C across diverse PC cell lines. Treatment with VIC-1911, even at nanomolar concentrations, substantially inhibits the growth of both androgen receptor (AR)-positive and AR-negative PC cells. VIC-1911 triggers mitotic failure, induces DNA double-strand breaks (DSBs), and activates the p53 pathway, halting cell division and inducing cell death. Notably, VIC-1911 showed synergistic effects in inhibiting PC cell growth in vitro and xenograft tumor growth in vivo with poly (ADP-ribose) polymerase inhibitors (PARPi), which have proven effective in PC with a deficiency in Homologous Recombination (HR) repair. Mechanistically, VIC-1911 disabled HR-mediated repair of DSBs in otherwise HR-proficient PC cells, leading to a “BRCAness” phenotype and pronounced accumulation of DNA damage and mitotic catastrophe. In summary, our study uncovers what we believe a novel mechanism to functional “BRCAness” by inducing mitotic arrest and highlights VIC-1911 as a promising therapeutic agent for advanced PC, either as a single agent or in combination, sensitizing HR-proficient tumors to PARP inhibitors.
Authors
Galina Gritsina, Sandip Kumar Rath, Hongshun Shi, Qi Chu, Wanqing Xie, Que Thanh Thanh Nguyen, Sambhavi Senthil, Thomas J. Myers, Mehmet A. Bilen, Sarah E. Fenton, Maha Hussain, David S. Yu, Jonathan C. Zhao, Jindan Yu
Abstract
Spinal muscular atrophy (SMA) is a devastating neuromuscular disorder caused by mutations in the survival motor neuron 1 (SMN1) gene leading to decreased SMN protein levels and motor neuron dysfunction. SMN-restoring therapies offer clinical benefit, but the downstream molecular consequences of SMN reduction remain incompletely understood. SMN deficiency resulted in downregulation of kinesin heavy chain isoform 5A (KIF5A) in human neurons and in a mouse model of SMA. SMN associated with KIF5A mRNA and contributed to its stability. Reduced SMN levels impaired axon regeneration, which was rescued by KIF5A overexpression. Because KIF5A has also been connected to ALS, these findings provide evidence of a molecular link between SMA and ALS pathophysiology, highlighting KIF5A as an SMN regulated factor. Our findings suggest SMN-independent interventions targeting KIF5A could represent a complementary therapeutic approach for SMA and other motor neuron diseases.
Authors
Tetsuya Akiyama, Yi Zeng, Caiwei Guo, Olivia Gautier, Lauren Koepke, Heankel Lyons, Elana Molotsky, Juliane S. Bombosch, Odilia Sianto, Jay P. Ross, Phuong Hoang, Luke Zhao, Cole Spencer, Charlotte J. Sumner, Michelle Monje, John W. Day, Aaron D. Gitler
Abstract
A large inter-individual variability in weight loss outcomes following bariatric surgery is reported. To ensure optimal patient management, it is crucial to accurately identify those most likely to benefit from the intervention. Since genetic variants largely contribute to surgery response, polygenic scores (PGS) derived from genome-wide association studies (GWAS) could constitute valuable tools for clinical decision making. We developed and evaluated PGS to predict the weight loss response in 540 patients with body mass index (BMI) ≥35kg/m2 who underwent biliopancreatic diversion with duodenal switch. Summary statistics derived from BMI-derived GWAS, together with summary statistics from previously published GWAS of BMI and adiposity features, were used to construct, evaluate, and benchmark weight-loss PGS. The full-adjusted BMI PGS model built in the entire cohort explained 39.6% of the mean-over-time excessive body weight loss (%EBWL), while the BMI-PGS built in the training dataset explained 38.9%. All benchmarked PGS based on BMI showed a significant relationship with mean-over-time %EBWL. These findings highlight the potential of BMI PGS in predicting weight loss after bariatric surgery and support their use as promising tools to improve the effectiveness of future anti-obesity treatments. Funding: Canadian Institutes of Health Research (PJT-168876).
Authors
Bastien Vallée Marcotte, Juan de Toro-Martín, André Tchernof, Louis Pérusse, Simon Marceau, Marie-Claude Vohl
Abstract
Cutaneous radiation injury is an unintended consequence of radiotherapy for many common cancers and can progress to debilitating radiation-induced skin fibrosis (RISF). Existing radiation injury models do not fully capture the skin toxicities observed in patients, contributing to the lack of efficacious therapies to mitigate RISF. To address this, we developed an ex vivo human skin model that recapitulates the temporal radiation injury and RISF response. Human skin explants (N=12) subjected to ionizing radiation demonstrated DNA double-strand breaks and robust p53-driven transcriptional programming of cell cycle arrest, apoptosis, and senescence compared to non-irradiated controls. Irradiated skin also exhibited induction of pro-inflammatory cytokines, epithelial-mesenchymal transition, pro-fibrotic TGF-beta1 (TGFB1)-mediated signaling, and thickened collagen over time. P53 regulators murine double minute 2 (MDM2) and microRNA (miR)-34a were induced post-irradiation and may be leveraged to modulate injury response. Notably, RNA-sequencing of breast skin from mastectomy patients post-radiotherapy showed similar p53, inflammatory, and TGFB1 signatures as the ex vivo model, supporting its translational relevance. Together, this model provides a platform for identifying biomarkers and testing therapies to prevent or mitigate cutaneous radiation toxicities. Targeting the dynamic p53-driven pro-fibrotic radiation response represents a new therapeutic avenue to improve post-radiotherapy quality of life for cancer survivors.
Authors
Caroline Dodson, Sophie M. Bilik, Gabrielle DiBartolomeo, Hannah Pachalis, Lindsey G. Siegfried, Jordan A. K. Johnson, Seth R. Thaller, Irena Pastar, Marjana Tomic-Canic, Anthony J. Griswold, Rivka C. Stone
No posts were found with this tag.